Note
Go to the end to download the full example code.
Energy and forces¶
Single-point evaluation of a bulk silicon cell with the smallest PET-MAD
model (pet-mad-xs). Two atoms are displaced off their equilibrium
positions so the predicted forces are non-trivial; we then attach a
UPETCalculator, read energy and forces via
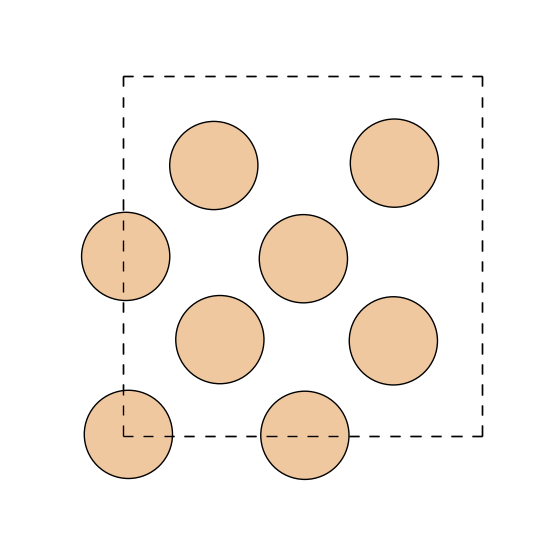
the standard ASE API, and render the unit cell with the forces drawn as
arrows on top of the atoms.
/home/runner/work/upet/upet/.tox/docs/lib/python3.13/site-packages/metatomic_ase/_neighbors.py:78: UserWarning: `compute_requested_neighbors_from_options` is deprecated and will be removed in a future version. Please use `neighbor_lists_for_model` to get the calculators and call them directly.
vesin.metatomic.compute_requested_neighbors_from_options(
Number of atoms: 8
Total energy: -46.7452 eV
Energy per atom: -5.8432 eV
Max |force| component: 1.7334e+00 eV/Å
Stress (Voigt, eV/ų): [-0.01 -0.0081 -0.0085 -0.0002 0.0159 0.0047]
import matplotlib.pyplot as plt
import numpy as np
from ase.build import bulk
from ase.visualize.plot import plot_atoms
from upet.calculator import UPETCalculator
atoms = bulk("Si", cubic=True, a=5.43, crystalstructure="diamond")
# perturb every atom by a small random displacement so the predicted
# forces on every site are non-zero and of comparable magnitude
atoms.rattle(0.05, seed=0) # ASE's built-in random displacement method
calculator = UPETCalculator(model="pet-mad-xs", version="1.5.0", device="cpu")
atoms.calc = calculator
energy = atoms.get_potential_energy()
forces = atoms.get_forces()
stress = atoms.get_stress()
print(f"Number of atoms: {len(atoms)}")
print(f"Total energy: {energy:.4f} eV")
print(f"Energy per atom: {energy / len(atoms):.4f} eV")
print(f"Max |force| component: {np.abs(forces).max():.4e} eV/Å")
print(f"Stress (Voigt, eV/ų): {np.array2string(stress, precision=4)}")
# Visualize the cell projected along z and overlay the in-plane force
# components as red arrows. A matplotlib quiver key acts as a scale bar.
fig, ax = plt.subplots(figsize=(5.5, 5.5))
plot_atoms(atoms, ax, radii=0.6, show_unit_cell=2)
plt.show()
Total running time of the script: (0 minutes 9.643 seconds)