Note
Go to the end to download the full example code.
NPT molecular dynamics¶
Isobaric–isothermal (constant pressure, constant temperature) MD of bulk
silicon with the ase.md.npt.NPT integrator (Parrinello–Rahman
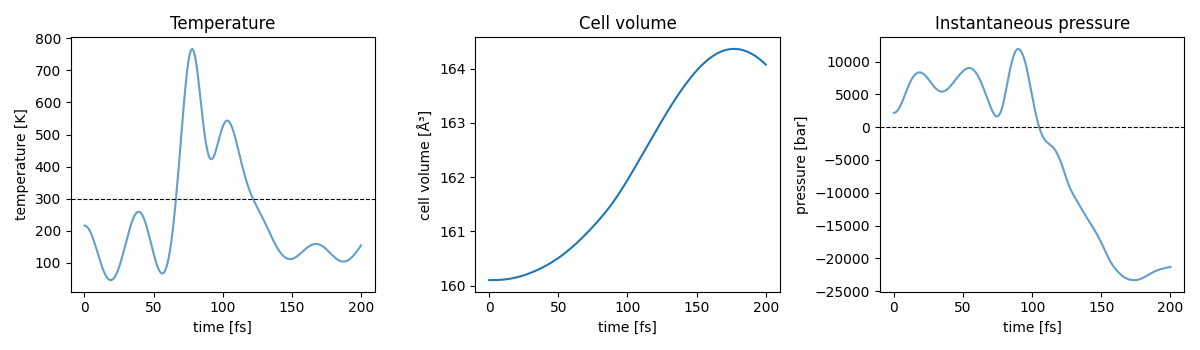
barostat coupled to a Nosé–Hoover thermostat) at 300 K and 1 bar. Both
the cell and the atomic coordinates evolve; we monitor temperature, cell
volume and instantaneous pressure.
/home/runner/work/upet/upet/examples/1-ase-simulations/plot_md_npt.py:38: FutureWarning: NPT thermostat has been moved/renamed to ase.md.melchionna.MelchionnaNPT. Please use this class instead (or a newer NPT thermostat!)
dyn = NPT(
WARNING: NPT: Setting the center-of-mass momentum to zero (was -0.860604 -0.621768 -0.85177)
/home/runner/work/upet/upet/.tox/docs/lib/python3.13/site-packages/metatomic_ase/_neighbors.py:78: UserWarning: `compute_requested_neighbors_from_options` is deprecated and will be removed in a future version. Please use `neighbor_lists_for_model` to get the calculators and call them directly.
vesin.metatomic.compute_requested_neighbors_from_options(
import matplotlib.pyplot as plt
import numpy as np
from ase import units
from ase.build import bulk
from ase.md.npt import NPT
from ase.md.velocitydistribution import MaxwellBoltzmannDistribution
from upet.calculator import UPETCalculator
T_TARGET = 300.0 # K
P_TARGET_BAR = 1.0
bar = 1e-4 * units.GPa # ASE uses eV/ų for stress
p_target = P_TARGET_BAR * bar
atoms = bulk("Si", cubic=True, a=5.43, crystalstructure="diamond")
atoms.calc = UPETCalculator(model="pet-mad-xs", version="1.5.0", device="cpu")
MaxwellBoltzmannDistribution(
atoms, temperature_K=T_TARGET, rng=np.random.default_rng(0)
)
ttime = 25.0 * units.fs
ptime = 100.0 * units.fs
bulk_modulus_si = 100.0 * units.GPa # Silicon ≈ 98 GPa
dyn = NPT(
atoms,
timestep=1.0 * units.fs,
temperature_K=T_TARGET,
externalstress=p_target,
ttime=ttime,
pfactor=(ptime**2) * bulk_modulus_si,
)
n_steps = 200
times, temps, volumes, pressures = [], [], [], [] # type: ignore
def log():
times.append(len(times) * 1.0)
temps.append(atoms.get_temperature())
volumes.append(atoms.get_volume())
# ASE stress is Voigt (xx, yy, zz, yz, xz, xy); pressure = -trace/3
s = atoms.get_stress()
pressures.append(-(s[0] + s[1] + s[2]) / 3.0 / bar)
log()
for _ in range(n_steps):
dyn.run(1)
log()
times = np.asarray(times)
temps = np.asarray(temps)
volumes = np.asarray(volumes)
pressures = np.asarray(pressures)
fig, axes = plt.subplots(1, 3, figsize=(12, 3.5))
axes[0].plot(times, temps, alpha=0.7)
axes[0].axhline(T_TARGET, color="k", ls="--", lw=0.8)
axes[0].set_xlabel("time [fs]")
axes[0].set_ylabel("temperature [K]")
axes[0].set_title("Temperature")
axes[1].plot(times, volumes)
axes[1].set_xlabel("time [fs]")
axes[1].set_ylabel("cell volume [ų]")
axes[1].set_title("Cell volume")
axes[2].plot(times, pressures, alpha=0.7)
axes[2].axhline(P_TARGET_BAR, color="k", ls="--", lw=0.8)
axes[2].set_xlabel("time [fs]")
axes[2].set_ylabel("pressure [bar]")
axes[2].set_title("Instantaneous pressure")
fig.tight_layout()
plt.show()