Note
Go to the end to download the full example code.
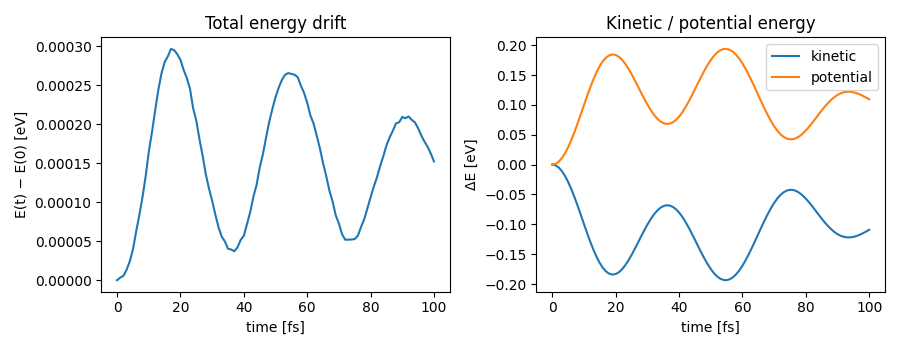
NVE molecular dynamics¶
Microcanonical (constant energy) molecular dynamics of bulk silicon using
ase.md.verlet.VelocityVerlet. Velocities are initialized from
a Maxwell–Boltzmann distribution at 300 K; total energy should stay flat
up to integration noise.
/home/runner/work/upet/upet/.tox/docs/lib/python3.13/site-packages/metatomic_ase/_neighbors.py:78: UserWarning: `compute_requested_neighbors_from_options` is deprecated and will be removed in a future version. Please use `neighbor_lists_for_model` to get the calculators and call them directly.
vesin.metatomic.compute_requested_neighbors_from_options(
import matplotlib.pyplot as plt
import numpy as np
from ase import units
from ase.build import bulk
from ase.md.velocitydistribution import MaxwellBoltzmannDistribution
from ase.md.verlet import VelocityVerlet
from upet.calculator import UPETCalculator
atoms = bulk("Si", cubic=True, a=5.43, crystalstructure="diamond")
atoms.calc = UPETCalculator(model="pet-mad-xs", version="1.5.0", device="cpu")
MaxwellBoltzmannDistribution(atoms, temperature_K=300, rng=np.random.default_rng(0))
dyn = VelocityVerlet(atoms, timestep=1.0 * units.fs)
n_steps = 100
times, e_tot, e_kin, e_pot = [], [], [], [] # type: ignore
def log():
times.append(len(times) * 1.0) # fs
ekin = atoms.get_kinetic_energy()
epot = atoms.get_potential_energy()
e_kin.append(ekin)
e_pot.append(epot)
e_tot.append(ekin + epot)
log()
for _ in range(n_steps):
dyn.run(1)
log()
times = np.asarray(times)
e_tot = np.asarray(e_tot)
e_kin = np.asarray(e_kin)
e_pot = np.asarray(e_pot)
fig, (ax_tot, ax_split) = plt.subplots(1, 2, figsize=(9, 3.5))
ax_tot.plot(times, e_tot - e_tot[0])
ax_tot.set_xlabel("time [fs]")
ax_tot.set_ylabel("E(t) − E(0) [eV]")
ax_tot.set_title("Total energy drift")
ax_split.plot(times, e_kin - e_kin[0], label="kinetic")
ax_split.plot(times, e_pot - e_pot[0], label="potential")
ax_split.set_xlabel("time [fs]")
ax_split.set_ylabel("ΔE [eV]")
ax_split.set_title("Kinetic / potential energy")
ax_split.legend()
fig.tight_layout()
plt.show()